This project focuses on building an automated image processing pipeline to process, align, and morph consecutive slices of tissue samples from Whole Slide Images (WSI). By correcting structural deformations introduced during physical tissue sectioning and scanning, the pipeline is able to construct smooth intermediate representations (morphs) between sequential tissue layers.
This approach helps in visualizing gradual morphological changes in biological tissues in 3D-like continuity.
| Authors | Jedrzej Golaszewski |
|---|---|
| Title | MiMorph |
| Status | Completed |
| Date | Sep 2, 2025 |
| Tech stack | OpenSlide, scikit-image, OpenCV, SimpleITK, PyWavelets, Optuna, NumPy |
| Links |
Pipeline Architecture
The pipeline is split into four distinct stages, moving from raw slide extraction to the final morphed outputs.
1. Tissue Preparation
The first step involves isolating individual tissue regions from high-resolution WSI (Whole Slide Images) scans. Biological samples are often scanned at gigapixel resolutions and saved in specialized formats like .mrxs. Before any image processing can begin, we must handle these massive files, typically using the OpenSlide library to extract data at a manageable Level of Detail (LOD).
At this raw stage, a single digital slide often contains multiple tissue sections scattered across the glass, alongside bright backgrounds, dust, and sometimes marker ink.

To prepare these messy, unstandardized samples for precise mathematical alignment, the pipeline executes a series of automated cleaning and extraction steps:
Color Deconvolution: Because tissues are typically stained with Hematoxylin and Eosin (H&E) for microscopic examination, the images are captured in standard RGB color space. However, RGB is not ideal for distinguishing biological structures from the background glass. Using
scikit-image, we apply color deconvolution to transform the image into the HED (Hematoxylin-Eosin-DAB) color space. This process mathematically separates the specific purple and pink stain channels, allowing us to isolate the actual tissue matter from background artifacts.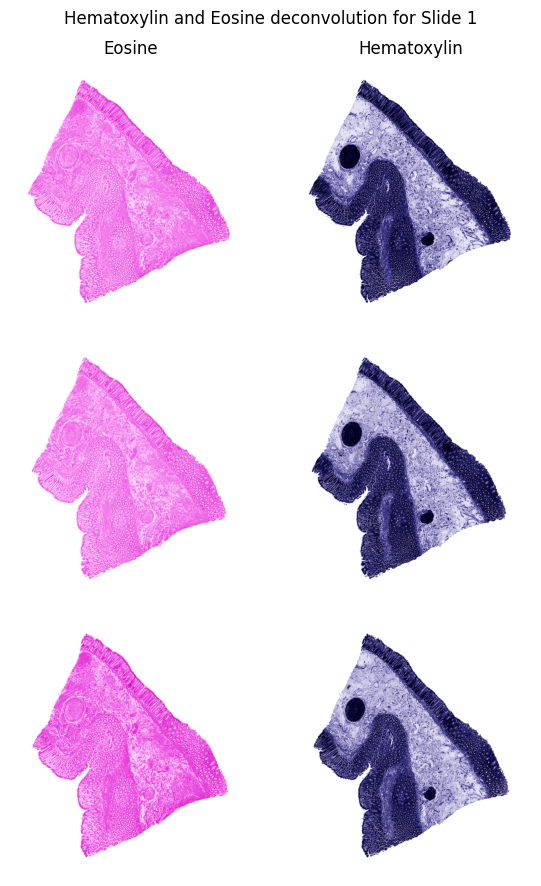
Extraction & Masking: Once the tissue structures are clearly segmented in the HED color space, the pipeline uses
OpenCVto identify the exact physical boundaries of each slice. We apply binary thresholding to create a stark contrast map, followed by contour detection algorithms to trace the outer edges of the tissues. By calculating bounding boxes around these detected contours, we can automatically crop each individual slice and generate crisp binary masks.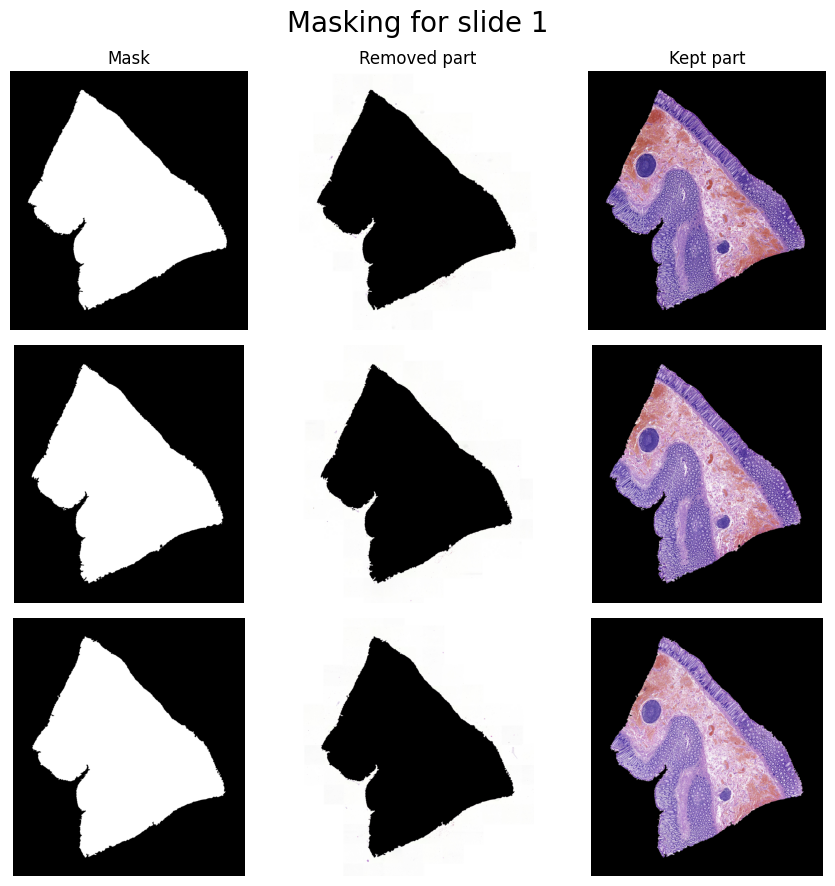
Standardization: The final result of this preparation phase is a structured dataset. What started as a single, noisy slide is now an organized collection of standardized, perfectly cropped tissue images alongside their corresponding masks. This guarantees that the downstream alignment algorithms process only the relevant biological data, free from background interference.
2. Affine Registration
Physical sectioning often causes the tissue slices to rotate, shift, or scale differently on the glass slides.
- Global Alignment: To correct global misalignments, we apply a multi-step Affine Registration process.
- Wavelet Pyramids: The process leverages a Bior6.8 wavelet transform pyramid to perform coarse-to-fine alignment, ensuring rapid and robust matching of structural features before moving on to complex warp techniques.
The result of this stage is a globally aligned stack of images, as demonstrated below:
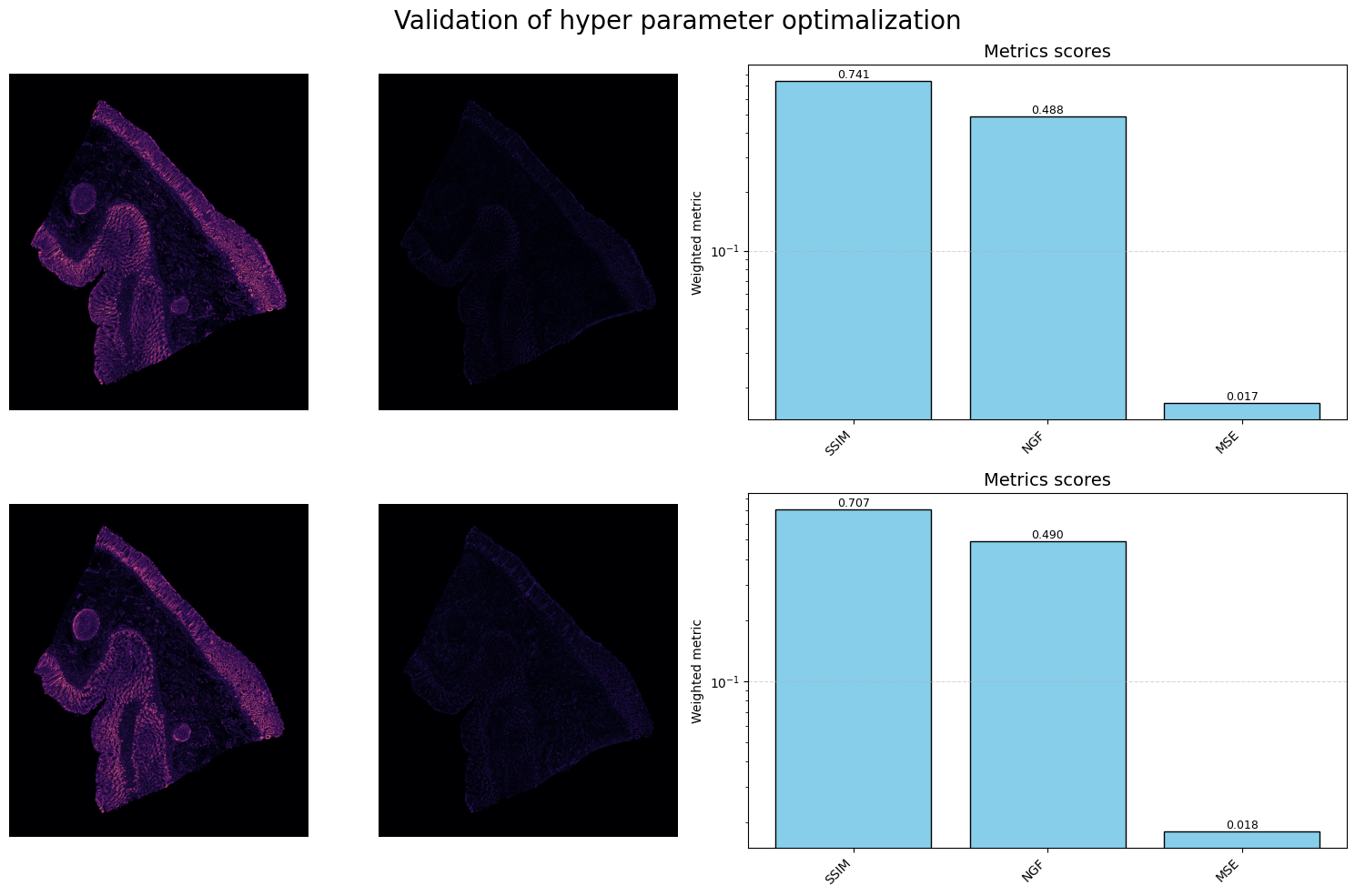
3. Non-Affine Registration
While affine transformations correct global shifts, tissues also undergo localized stretching, tearing, and non-linear deformations during the physical cutting and mounting process.
- Mattes Mutual Information (MMI): We use MMI-based non-affine (B-spline/deformable) registration via SimpleITK to correct these localized distortions.
- Hyperparameter Optimization: Because non-affine alignment is highly sensitive to parameter tuning, we integrated Optuna to optimize the registration hyperparameters, maximizing structural preservation and edge alignment.
This step generates a localized deformation field that shifts internal structures into their proper anatomical alignment without affecting the global orientation:
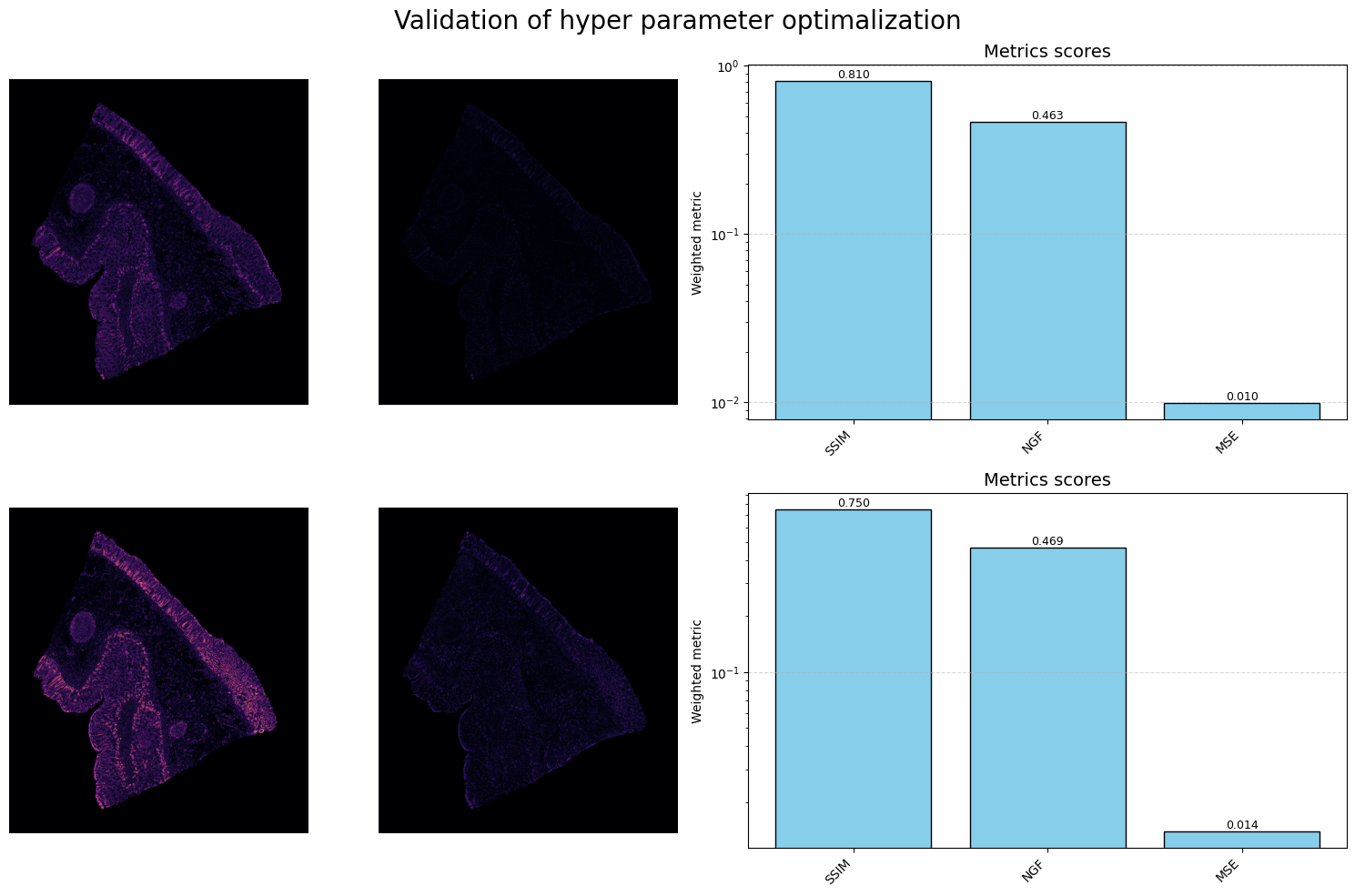
4. Tissue Morphing
Once consecutive tissue layers are accurately aligned globally and locally, the final stage generates the intermediate tissue states.
- Optical Flow: By computing the optical flow between an aligned tissue slice and its subsequent slice, the pipeline generates smooth visual transitions (warping).
- Blending: The warped images are alpha-blended to simulate the continuous biological structures that exist between the physical cuts.
The final output is a seamless morphing sequence that bridges the physical gap between slices:
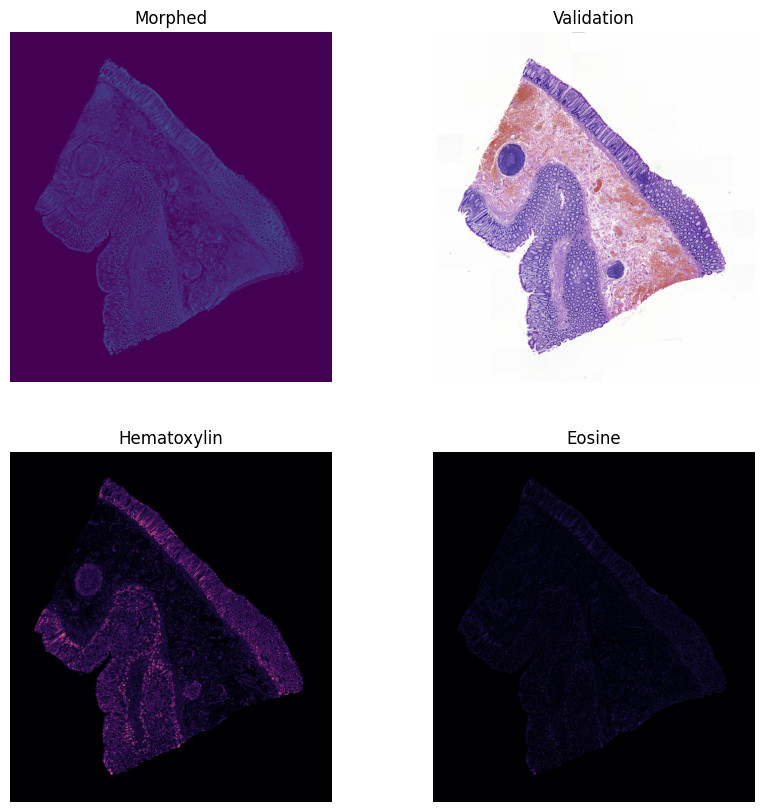

Conclusions & Future Work
While the pipeline successfully generates morphed transitions, the current effectiveness is constrained by a few factors:
- Alignment Tuning: The non-affine alignment was originally tuned for consecutive slices. When applied to morphing (which sometimes attempts to bridge larger gaps or every second slice), the effectiveness drops.
- Future Directions: Future iterations will focus on optimizing alignment parameters specifically for non-consecutive sequences. Additionally, with access to larger annotated datasets, the pipeline could benefit significantly from integrating Deep Learning and AI-based registration/morphing techniques to handle complex tearing and artifacts better than traditional optical flow.
Tools and Libraries used: OpenSlide, scikit-image, OpenCV, SimpleITK, PyWavelets, Optuna, NumPy.